
Research Interests
Bioinformatics Bioinformatics Software Biological Evolutionary Biology Genetics Genomics Medical GeneticsTeaching Strengths
Dr Yassine Souilmi
NHMRC Grant-Funded Research Fellow
School of Biological Sciences
College of Science
Eligible to supervise Masters and PhD - email supervisor to discuss availability.
I am the Group Leader for Genomics and Bioinformatics at the Australian Centre for Ancient DNA (ACAD), and an Evolutionary Medicine Research Fellow at the Black Ochre Data Labs (BODL) — a joint initiative between the Kids Research Institute Australia and the Australian National University. My work uses ancient and modern DNA, large-scale computation, and reproducible bioinformatics to understand how past events shape present-day human and animal health. My current programme runs along two concurrent tracks: Indigenous genomics conducted under Indigenous governance and leadership at BODL, and dingo genomics and conservation in collaboration with Indigenous communities, government agencies, and industry.
Research impact at a glance
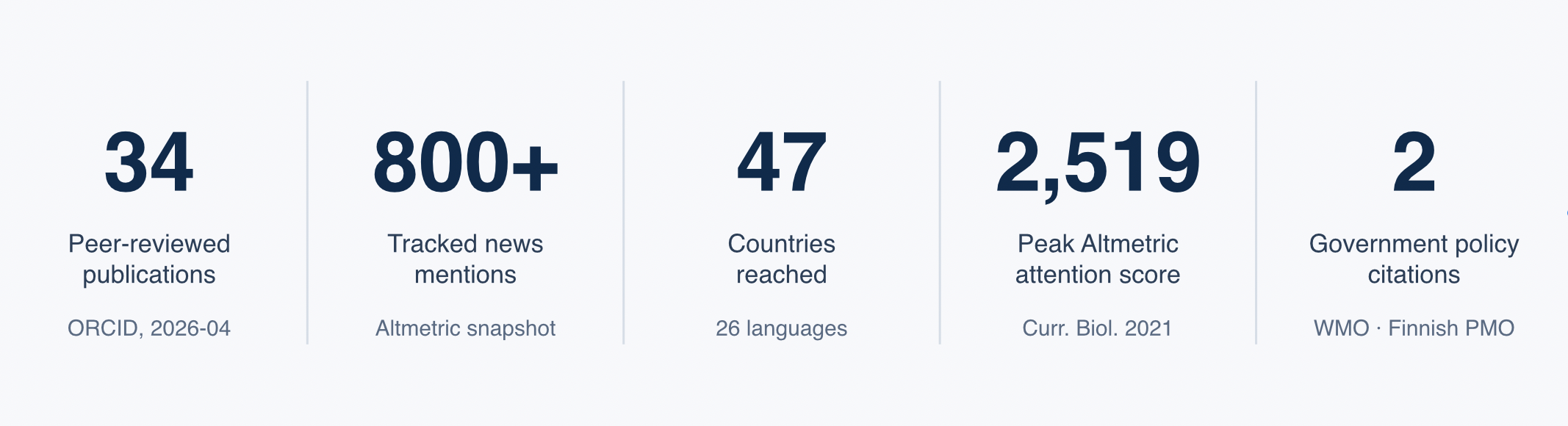
A short summary of where the work has landed beyond the academic literature; numbers draw from an Altmetric snapshot dated 2026-04-01.
- 34 peer-reviewed publications — live record on ORCID and Google Scholar.
- 800+ tracked news mentions across 47 countries and 26 languages.
- Peak Altmetric Attention Score 2,519 — Souilmi et al., Current Biology, 2021 (ancient coronavirus epidemic in East Asia).
- Two government / intergovernmental policy citations: World Meteorological Organization, 2024 ozone-science assessment; Finnish Prime Minister's Office, COVID-19 research review (2020).
- Tier-1 outlets covering my work: The New York Times, The Guardian, CNN, Smithsonian Magazine, National Geographic, New Scientist, Der Spiegel, Cosmos Magazine.
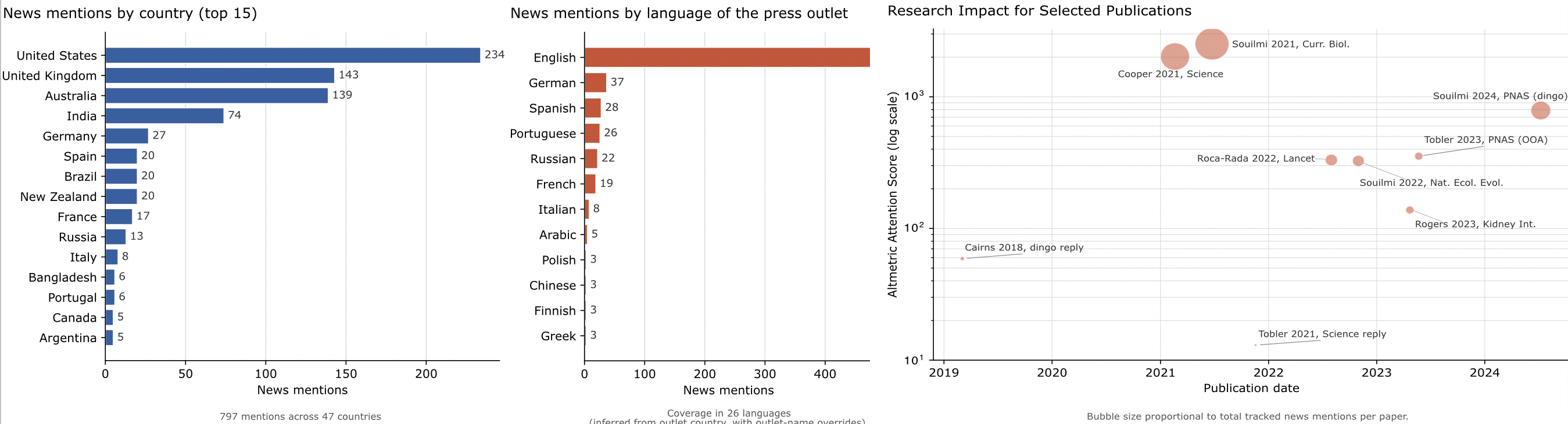
Last panel (rightmost) represents the engagement over time, sized by news mentions for a select number of manuscripts. (Altmetric, snapshot 2026-04-01).
Research themes
Evolutionary Medicine

Past selection has shaped present-day disease risk. In Souilmi et al., Current Biology (2021) we identified an ancient viral epidemic involving host coronavirus-interacting genes more than 20,000 years ago in East Asia, surfacing candidate genes relevant to modern coronaviruses; the paper remains the most publicly engaged piece in my catalogue (peak Altmetric 2,519, cited in the Finnish Prime Minister's Office COVID-19 research review). With Tobler and colleagues we extended this lens to the out-of-Africa expansion: genetic selection and climatic factors jointly shaped human dispersal (PNAS, 2023). I also led the demonstration that admixture has obscured signals of historical hard sweeps in humans (Nature Ecology & Evolution, 2022) — a methodological correction with implications for any selection scan that ignores admixture history.
Functional and clinical-translational work brings these signals into disease biology directly. With Sinniah and colleagues we showed in Nucleic Acids Research (2025) that conserved facultative heterochromatin marks regulatory sequences underpinning cell identity and disease — bridging population-scale evolutionary signal with mechanistic disease genomics. With Rogers and colleagues we characterised the impact of TNFAIP3 variation on NF-κB activation in acute kidney injury (Kidney International, 2023). And in a clinical-paleogenomics case study, the team integrated morphology, osteology, and ancient genomics to diagnose a 1,000-year-old case of Klinefelter's syndrome (The Lancet, 2022).
My current focus in this theme is Indigenous genomics, conducted under Indigenous governance and leadership. At BODL, I contribute genomics and bioinformatics expertise to community-led research using the largest cohort of Indigenous genomes in Australia, characterising the genetic determinants of Type 2 Diabetes in Aboriginal South Australians. The programme is supported by the MRFF Genomics Health Futures Mission grant "Pathways to benefit for Indigenous Australians in Genomic Medicine" (MRF2016124, A$4.99M), complemented by an MRFF-funded long-read nanopore sequencing platform for Indigenous genomics (MRF2016008, A$0.99M), and by the Australian Academy of Science Theo Murphy Initiative "Indigenous Genomics and Responsible Research" MOOC. Earlier collaborative work in this space includes Equitable Expanded Carrier Screening Needs Indigenous Clinical and Population Genomic Data (American Journal of Human Genetics, 2020) and the development of a draft Australian reference pangenome with the National Centre for Indigenous Genomics. Across all of it, the principle is the same: genomic methods are tools placed in service of Indigenous-led research priorities and community benefit.
Dingo Genomics & Conservation
Dingoes are an iconic, ecologically and culturally important free-living canid, and a focal organism for my conservation genomics work. In Souilmi et al., PNAS (2024), our team used ancient genomes to reveal over two thousand years of dingo population structure, providing a pre-Colonial baseline against which present-day admixture with domestic dogs can be quantified — directly informing dingo management and policy. The paper drew a fresh wave of tier-1 international coverage three years after the data were collected, and the 2026 commentary on gene-edited "super-quolls" (The Age, SMH, Brisbane Times, WA Today) picked up the same conservation-genomics thread. Earlier work in Zootaxa (2019) contributed to the long-running debate on dingo taxonomy.

On the methods side, our diploid dual canine assemblies (NAR Genomics and Bioinformatics, 2026) revealed extensive allelic heterogeneity and a telocentric chromosomal architecture — improving the genomic resources that underpin both wild- and companion-canid genetics. I also advise the Centre for Invasive Species Solutions and the South Australian Department of Primary Industries and Regions (PIRSA) on applied conservation and invasive-species genomics.
Bioinformatics, Methods & Cloud Infrastructure
Reproducibility and infrastructure underpin every theme above. I have worked with cloud-based scientific computing since 2012, when production-scale genomics on commercial cloud was still fringe. At Harvard Medical School I helped build the COSMOS distributed-pipeline framework and its bioinformatics pipelines (GenomeKey, PV-Key, and the multi-cloud variant MC-GenomeKey); these were later deployed at Invitae in clinical production, processing hundreds of thousands of patient samples — an early concrete demonstration that cloud-deployed genomic workflows could meet diagnostic-grade reliability.

Attribution-NonCommercial-NoDerivatives License 4.0
(CC BY).
At Adelaide I have played a key role in institutional adoption of AWS through the RONIN research-cloud platform, and have contributed open methods that the field now uses: the Genozip universal genomic-data compressor (Bioinformatics, 2021); systematic benchmarks of ancient-DNA read mapping (Briefings in Bioinformatics, 2021); the taxonomicfiltering Kraken2-based pre-mapping workflow that cuts ancient-DNA mapping runtime by up to ~94% (Briefings in Bioinformatics, 2024); and consensus recommendations for reproducible paleogenomic data analyses (American Journal of Human Genetics, 2025). My PhD student Shyamsundar Ravishankar won the AGTA 2024 best oral student presentation award for cloud-backed ancient-DNA workflows built on this stack.
Selected collaborative work in population genomics & deep history
Beyond the themes above, ongoing collaborations have reconstructed deep regional population histories: the genetic history of Portugal over the past 5,000 years (Genome Biology, 2025); Neolithic-to-Bronze-Age genetic transitions at Mas d'en Boixos in Catalonia (iScience, 2025); the early peopling of the Southern Cone of South America from ancient mitochondrial genomes (iScience, 2021); and the global environmental crisis 42,000 years ago (Science, 2021), since cited in the World Meteorological Organization's 2024 ozone-science assessment.
Funding and collaboration
I am currently a named investigator or substantive contributor on competitive grants totalling ~A$13.76M, including the MRFF Genomics Health Futures Mission (Indigenous Genomic Medicine; long-read nanopore platform), the MRFF Early-to-Mid-Career Researchers scheme (complete genomics, with the Garvan Institute / UNSW), the ARC Australian Laureate Fellowship programme, and the Australian Academy of Science Theo Murphy Initiative. I welcome collaborations in evolutionary genomics, ancient DNA, Indigenous-led genomics, conservation genomics, and reproducible bioinformatics, and I supervise Masters and PhD students in these areas.
Find me online: ORCID · Google Scholar · GitHub · The Conversation · ysouilmi.com
| Date | Position | Institution name |
|---|---|---|
| 2016 - ongoing | ARC Postdoctoral Researcher | University of Adelaide |
| 2014 - 2015 | Research and Teaching Assistant | Harvard University |
| 2014 - 2014 | Teaching Assistant | Harvard University |
| 2013 - 2013 | Adjunct Bioinformatics Instructor | Mohammed V University of Rabat |
| Date | Type | Title | Institution Name | Country | Amount |
|---|---|---|---|---|---|
| 2013 | Scholarship | Fulbright Joint Supervision Doctoral Grant | Harvard University | United States | - |
| Language | Competency |
|---|---|
| Arabic | Can read, write, speak, understand spoken and peer review |
| English | Can read, write, speak, understand spoken and peer review |
| French | Can read, write, speak, understand spoken and peer review |
| Spanish; Castilian | Can read and understand spoken |
| Date | Institution name | Country | Title |
|---|---|---|---|
| 2012 - 2016 | Mohammed V University | Morocco | PhD |
| 2008 - 2011 | Mohammed V University | Morocco | MS |
| 2004 - 2008 | Mohammed V University | Morocco | BS |
| Year | Citation |
|---|---|
| 2026 | Kidd, J. M., Souilmi, Y., Rosen, B. D., Khan, R., Weisz, D., Dudchenko, O., . . . Ballard, J. W. O. (2026). Diploid dual assemblies reveal the telocentric structure and extensive allelic heterogeneity of canine genomes. NAR Genomics and Bioinformatics, 8(2), lqag035-1-lqag035-12. |
| 2026 | Ravishankar, S., Nguyen, N. C., Taufik, L., Michielsen, N. M., Bergström, A., Tobler, R., . . . Souilmi, Y. (2026). Paleogenomics‐Informed Inferences of European Dog Admixture Enables Scalable Dingo Conservation. Conservation Letters, 19(3). |
| 2025 | Ravishankar, S., Perez, V., Davidson, R., Roca-Rada, X., Lan, D., Souilmi, Y., & Llamas, B. (2025). Filtering out the noise: metagenomic classifiers optimize ancient DNA mapping. Briefings in Bioinformatics, 26(1), bbae646-1-bbae646-12. Scopus5 WoS4 Europe PMC3 |
| 2025 | Roca-Rada, X., Cuesta-Aguirre, D. R., Vinueza-Espinosa, D. C., Davidson, R., Ravishankar, S., Taufik, L., . . . Santos, C. (2025). Genetic transitions in the Neolithic and Bronze Age at Mas d’en Boixos (Catalonia, Spain). iScience, 28(7), 112871-1-112871-11. Scopus2 WoS2 Europe PMC1 |
| 2025 | Davidson, R., Ravishankar, S., Souilmi, Y., Roca-Rada, X., Sobek, C., Taufik, L., . . . Pérez, V. (2025). The necessity for authentication of ancient DNA from archaeological artefacts. Journal of Archaeological Science, 181, 106317-1-106317-9. |
| 2025 | Roca-Rada, X., Davidson, R., Williams, M. P., Villalba-Mouco, V., Carvalho, A. F., Ravishankar, S., . . . Teixeira, J. C. (2025). The genetic history of Portugal over the past 5,000 years.. Genome Biology, 26(1), 248-1-248-39. Scopus5 WoS6 Europe PMC1 |
| 2025 | Souilmi, Y., Oliva, A., Davidson, R., Williams, M. P., Ravishankar, S., Roca-Rada, X., . . . Llamas, B. (2025). Lessons learned: Recommendations for reproducible paleogenomic data analyses.. Am J Hum Genet, 112(12), 2830-2841. |
| 2025 | Sinniah, E., Mizikovsky, D., Shim, W. J., Yeung Chow, C. S., Souilmi, Y., Cheng, F. -F., . . . Palpant, N. J. (2025). Conserved facultative heterochromatin across cell types identify regulatory sequences underpinning cell identity and disease. Nucleic Acids Research (NAR), 53(20), gkaf971-1-gkaf971-23. |
| 2024 | Souilmi, Y., Wasef, S., Williams, M. P., Conroy, G., Bar, I., Bover, P., . . . Mitchell, K. J. (2024). Ancient genomes reveal over two thousand years of dingo population structure. Proceedings of the National Academy of Sciences, 121(30), e2407584121-1-e2407584121-12. Scopus9 WoS8 Europe PMC6 |
| 2023 | Rogers, N. M., Zammit, N., Nguyen-Ngo, D., Souilmi, Y., Minhas, N., Meijles, D. N., . . . Grey, S. T. (2023). The impact of the cytoplasmic ubiquitin ligase TNFAIP3 gene variation on transcription factor NF-κB activation in acute kidney injury. Kidney international, 103(6), 1105-1119. Scopus15 WoS15 Europe PMC15 |
| 2023 | Tobler, R., Souilmi, Y., Huber, C. D., Bean, N., Turney, C. S. M., Grey, S. T., & Cooper, A. (2023). The role of genetic selection and climatic factors in the dispersal of anatomically modern humans out of Africa.. Proc Natl Acad Sci U S A, 120(22), e2213061120. Scopus22 WoS18 Europe PMC15 |
| 2022 | Lan, D., Purnomo, G., Tobler, R., Souilmi, Y., & Llamas, B. (2022). Genozip Dual-Coordinate VCF format enables efficient genomic analyses and alleviates liftover limitations. |
| 2022 | Roca-Rada, X., Tereso, S., Rohrlach, A. B., Brito, A., Williams, M. P., Umbelino, C., . . . Teixeira, J. C. (2022). A 1000-year-old case of Klinefelter's syndrome diagnosed by integrating morphology, osteology, and genetics.. Lancet (London, England), 400(10353), 691-692. Scopus12 WoS10 Europe PMC5 |
| 2022 | Souilmi, Y., Tobler, R., Johar, A., Williams, M., Grey, S. T., Schmidt, J., . . . Huber, C. D. (2022). Admixture has obscured signals of historical hard sweeps in humans. Nature Ecology & Evolution, 6(12), 2003-2015. Scopus22 WoS20 Europe PMC32 |
| 2021 | Oliva, A., Tobler, R., Cooper, A., Llamas, B., & Souilmi, Y. (2021). Systematic benchmark of ancient DNA read mapping. Briefings in bioinformatics, 22(5), 1-12. Scopus37 WoS34 Europe PMC34 |
| 2021 | Roca-Rada, X., Politis, G., Messineo, P. G., Scheifler, N., Scabuzzo, C., Gonzalez, M., . . . Fehren-Schmitz, L. (2021). Ancient mitochondrial genomes from the Argentinian Pampas inform the early peopling of the Southern Cone of South America. iScience, 24(6), 1-18. Scopus22 WoS23 Europe PMC9 |
| 2021 | Souilmi, Y., Lauterbur, M. E., Tobler, R., Huber, C. D., Johar, A. S., Moradi, S. V., . . . Enard, D. (2021). An ancient viral epidemic involving host coronavirus interacting genes more than 20,000 years ago in East Asia. Current Biology, 31(16), 3504-3514. Scopus65 WoS61 Europe PMC61 |
| 2021 | Oliva, A., Tobler, R., Llamas, B., & Souilmi, Y. (2021). BWA-mem is not the best aligner for ancient DNA short reads. Europe PMC2 |
| 2021 | Souilmi, Y., Lauterbur, M. E., Tobler, R., Huber, C. D., Johar, A. S., Moradi, S. V., . . . Enard, D. (2021). An ancient viral epidemic involving host coronavirus interacting genes more than 20,000 years ago in East Asia.. Curr Biol, 31(16), 3704. Scopus4 WoS2 Europe PMC11 |
| 2021 | Tobler, R., Souilmi, Y., Huber, C., Bean, N., Turney, C., Cooper, A., & Grey, S. (2021). Genetic and climatic factors in the dispersal of Anatomically Modern Humans Out of Africa. |
| 2021 | Cooper, A., Turney, C. S. M., Palmer, J., Hogg, A., McGlone, M., Wilmshurst, J., . . . Zech, R. (2021). Response to Comment on "A global environmental crisis 42,000 years ago". Science New York N Y, 374(6570), eabi9756. Scopus2 WoS2 Europe PMC2 |
| 2021 | Cooper, A., Turney, C. S. M., Palmer, J., Hogg, A., McGlone, M., Wilmshurst, J., . . . Zech, R. (2021). Response to comment on “A global environmental crisis 42,000 years ago”. Science, 374(6570), 3 pages. Scopus2 WoS1 |
| 2021 | Oliva, A., Tobler, R., Llamas, B., & Souilmi, Y. (2021). Additional evaluations show that specific BWA‐aln settings still outperform BWA‐mem for ancient DNA data alignment. Ecology and Evolution, 11(24), 18743-18748. Scopus18 WoS16 Europe PMC19 |
| 2021 | Cooper, A., Turney, C. S. M., Palmer, J., Hogg, A., McGlone, M., Wilmshurst, J., . . . Zech, R. (2021). A global environmental crisis 42,000 years ago. Science in China Series C: Life Sciences, 371(6531), 811-818. Scopus96 WoS86 Europe PMC23 |
| 2021 | Lan, D., Tobler, R., Souilmi, Y., & Llamas, B. (2021). Genozip - a universal extensible genomic data compressor. Bioinformatics, 37(16), 2225-2230. Scopus39 WoS32 Europe PMC26 |
| 2020 | Tobler, R., Johar, A., Huber, C., & Souilmi, Y. (2020). PolyLinkR: A linkage-sensitive gene set enrichment R package. |
| 2020 | Lan, D., Tobler, R., Souilmi, Y., & Llamas, B. (2020). genozip: a fast and efficient compression tool for VCF files. Bioinformatics, 36(13), 4091-4092. Scopus12 WoS10 Europe PMC13 |
| 2020 | Souilmi, Y., Tobler, R., Johar, A., Williams, M., Grey, S., Schmidt, J., . . . Cooper, A. (2020). Ancient human genomes reveal a hidden history of strong selection in Eurasia. Europe PMC1 |
| 2020 | Easteal, S., Arkell, R. M., Balboa, R. F., Bellingham, S. A., Brown, A. D., Calma, T., . . . Baynam, G. (2020). Equitable expanded carrier screening needs indigenous clinical and population genomic data. American Journal of Human Genetics, 107(2), 175-182. Scopus29 WoS27 Europe PMC25 |
| 2020 | Roca-Rada, X., Souilmi, Y., Teixeira, J. C., & Llamas, B. (2020). Ancient DNA studies in pre-Columbian Mesoamerica. Genes, 11(11), 1-19. Scopus8 WoS7 Europe PMC6 |
| 2020 | Souilmi, Y., Lauterbur, E., Tobler, R., Huber, C., Johar, A., & Enard, D. (2020). An ancient viral epidemic involving host coronavirus interacting genes more than 20,000 years ago in East Asia. Europe PMC3 |
| 2019 | Jackson, S. M., Fleming, P. J. S., Eldridge, M. D. B., Ingleby, S., Flannery, T., Johnson, R. N., . . . Helgen, K. M. (2019). The Dogma of Dingoes—Taxonomic status of the dingo: A reply to Smith et al.. Zootaxa, 4564(1), 198-212. Scopus43 WoS38 Europe PMC17 |
| 2019 | Llamas, B., Narzisi, G., Schneider, V., Audano, P., Biederstedt, E., Blauvelt, L., . . . Busby, B. (2019). A strategy for building and using a human reference pangenome. F1000Research, 8, 1751. Scopus4 Europe PMC8 |
| 2019 | Llamas, B., Narzisi, G., Schneider, V., Audano, P., Biederstedt, E., Blauvelt, L., . . . Busby, B. (2019). A strategy for building and using a human reference pangenome. F1000research, 8, 1751. Scopus11 Europe PMC1 |
| 2019 | Ahmed, A., Mpangase, P., Panji, S., Baichoo, S., Souilmi, Y., Fadlelmola, F., . . . Mulder, N. (2019). Organizing and running bioinformatics hackathons within Africa: The H3ABioNet cloud computing experience. AAS Open Res, 1, 9. |
| 2018 | Ahmed, A., Mpangase, P., Panji, S., Baichoo, S., Souilmi, Y., Fadlelmola, F., . . . Mulder, N. (2018). Organizing and running bioinformatics hackathons within Africa: The H3ABioNet cloud computing experience. |
| 2018 | Baichoo, S., Souilmi, Y., Panji, S., Botha, G., Meintjes, A., Hazelhurst, S., . . . Mulder, N. (2018). Developing reproducible bioinformatics analysis workflows for heterogeneous computing environments to support African genomics. BMC Bioinformatics, 19(1), 457-1-457-13. Scopus34 WoS32 Europe PMC31 |
| 2018 | Ahmed, A. E., Mpangase, P. T., Panji, S., Baichoo, S., Souilmi, Y., Fadlelmola, F. M., . . . Mulder, N. (2018). Organizing and running bioinformatics hackathons within Africa: The H3ABioNet cloud computing experience.. AAS open research, 1, 9. Europe PMC11 |
| 2017 | Elshazly, H., Souilmi, Y., Tonellato, P., Wall, D., & Abouelhoda, M. (2017). MC-GenomeKey: A multicloud system for the detection and annotation of genomic variants. BMC Bioinformatics, 18(1), 1-14. Scopus12 WoS11 Europe PMC5 |
| 2015 | Souilmi, Y., Allali, I., Badad, O., & Dwibedi, C. K. (2015). Highlights of the first ISCB Student Council Symposium in Africa 2015. F1000Research, 4, ISCB Comm J-569. Scopus5 Europe PMC10 |
| 2015 | Souilmi, Y., Jung, J. -Y., Lancaster, A., Gafni, E., Amzazi, S., Ghazal, H., . . . Tonellato, P. (2015). COSMOS: cloud enabled NGS analysis. BMC BIOINFORMATICS, 16(Suppl 2), 1 page. WoS1 Europe PMC1 |
| 2015 | Souilmi, Y., Lancaster, A., Jung, J., Rizzo, E., Hawkins, J., Powles, R., . . . Wall, D. (2015). Scalable and cost-effective NGS genotyping in the cloud. BMC Medical Genomics, 8(1), 64-1-64-9. Scopus19 WoS18 Europe PMC17 |
| 2014 | Gafni, E., Luquette, L. J., Lancaster, A. K., Hawkins, J. B., Jung, J. Y., Souilmi, Y., . . . Tonellato, P. J. (2014). COSMOS: Python library for massively parallel workflows. Bioinformatics Oxford England, 30(20), 2956-2958. Scopus17 WoS14 Europe PMC13 |
| - | Souilmi, Y. (n.d.). Supplemental data for: Ancient Genomes Reveal Over Two Thousand Years Of Dingo Population Structure. |
| Year | Citation |
|---|---|
| 2024 | Deshpande, A., Rogers, N., Grey, S., Souilmi, Y., & Li, J. (2024). INVESTIGATING THE ROLE OF TNFAIP3 VARIATION IN KIDNEY DISEASE. In NEPHROLOGY Vol. 29 (pp. 11). WILEY. |
| 2005 | Souilmi, Y., Knopp, R., & Caire, G. (2005). Code constructions for non-coherent <i>on-off</i> ultra-wideband systems. In 2005 IEEE INTERNATIONAL CONFERENCE ON ULTRA-WIDEBAND (ICU) (pp. 28-32). SWITZERLAND, Zurich: IEEE. DOI |
| 2004 | Knopp, R., & Souilmi, Y. (2004). Achievable rates for UWB peer-to-peer networks. In 2004 INTERNATIONAL ZURICH SEMINAR ON COMMUNICATIONS: ACCESS-TRANSMISSION-NETWORKING, PROCEEDINGS (pp. 82-85). SWITZERLAND, ETH, Zurich: IEEE. WoS8 |
| 2003 | Souilmi, Y., & Knopp, R. (2003). On the achievable rates of ultra-wideband systems in multipath fading environments. In 2003 IEEE INTERNATIONAL SYMPOSIUM ON INFORMATION THEORY - PROCEEDINGS (pp. 387). JAPAN, Yokohama: IEEE. |
| 2003 | Souilmi, Y., & Knopp, R. (2003). On the achievable rates of ultra-wideband PPM with non-coherent detection in multipath environments. In 2003 IEEE INTERNATIONAL CONFERENCE ON COMMUNICATIONS, VOLS 1-5 (pp. 3530-3534). AK, ANCHORAGE: IEEE. WoS25 |
| 2002 | Souilmi, Y., Rigazio, L., Nguyen, P., Kryze, D., & Junqua, J. C. (2002). Blind channel estimation based on speech correlation structure. In 2002 IEEE INTERNATIONAL CONFERENCE ON ACOUSTICS, SPEECH, AND SIGNAL PROCESSING, VOLS I-IV, PROCEEDINGS (pp. 393-396). FL, ORLANDO: IEEE. |
| Year | Citation |
|---|---|
| 2021 | Souilmi, Y., Lauterbur, E., Tobler, R., Huber, C., Johar, A. S., & Enard, D. (2021). An ancient coronavirus-like epidemic 25,000 years in ancestral populations from East Asia. Poster session presented at the meeting of AMERICAN JOURNAL OF PHYSICAL ANTHROPOLOGY. WILEY. |
| Year | Citation |
|---|---|
| - | Ravishankar, S., Nguyen, C. N., Michielsen, N., Bergström, A., Tobler, R., Fordham, D., . . . Souilmi, Y. (n.d.). Palaeogenomics-informed inferences of European dog admixture enables scalable dingo conservation. DOI |
| - | Souilmi, Y., & Tobler, R. (n.d.). Frequency data and SweepFinder2 output. DOI |
| - | Tobler, R., Souilmi, Y., & Huber, C. (n.d.). Supplemental Dataset S57. DOI |
| Year | Citation |
|---|---|
| 2026 | Ravishankar, S., Nguyen, N. C., Taufik, L., Michielsen, N. M., Bergström, A., Tobler, R., . . . Souilmi, Y. (2026). Palaeogenomics-informed inferences of European dog admixture enables scalable dingo conservation. DOI |
| 2024 | Roca-Rada, X., Davidson, R., Williams, M., Ravishankar, S., Collen, E., Haarkötter, C., . . . Teixeira, J. (2024). The genetic history of Portugal over the past 5,000 years. DOI |
I am currently a named investigator or substantive contributor on competitive grants totalling ~A$13.76M, including the MRFF Genomics Health Futures Mission (Indigenous Genomic Medicine; long-read nanopore platform), the MRFF Early-to-Mid-Career Researchers scheme (complete genomics, with the Garvan Institute / UNSW), the ARC Australian Laureate Fellowship programme, and the Australian Academy of Science Theo Murphy Initiative. I welcome collaborations in evolutionary genomics, ancient DNA, Indigenous-led genomics, conservation genomics, and reproducible bioinformatics, and I supervise Masters and PhD students in these areas.
I have an extensive bioinformatics teaching experience, having participated in teaching activities at the University of Mohamed the Vth and at Harvard University (BMI 714) where I designed, taught and graded bioinformatics practicums between 2014 and 2016. Furthermore, I have trained, organised and ran over 25 different bioinformatics workshops in Africa, the United States, and Australia covering a wide range of basic to advanced topics.
| Date | Role | Research Topic | Program | Degree Type | Student Load | Student Name |
|---|---|---|---|---|---|---|
| 2025 | Principal Supervisor | A Study of Ancient Pandemics in Human History Based on Virus Interacting Protein Genes | Master of Philosophy | Master | Full Time | Ms Huize Gao |
| 2025 | Principal Supervisor | A Study of Ancient Pandemics in Human History Based on Virus Interacting Protein Genes | Master of Philosophy | Master | Full Time | Ms Huize Gao |
| 2024 | Co-Supervisor | Genomic time travel: Unravelling the evolutionary history of fauna using paleogenomics | Doctor of Philosophy | Doctorate | Full Time | Mr Shyamsundar Ravishankar |
| 2024 | Co-Supervisor | Genomic time travel: Unravelling the evolutionary history of fauna using paleogenomics | Doctor of Philosophy | Doctorate | Full Time | Mr Shyamsundar Ravishankar |
| Date | Role | Research Topic | Program | Degree Type | Student Load | Student Name |
|---|---|---|---|---|---|---|
| 2021 - 2022 | Co-Supervisor | Advances in Genomic Data Compression | Doctor of Philosophy | Doctorate | Full Time | Mr Divon Mordechai Lan |
| 2019 - 2022 | Co-Supervisor | Human demographic insights from paleogenetic studies of American and Iberian skeletal remains | Doctor of Philosophy | Doctorate | Full Time | Mr Xavier Roca Rada |
| 2018 - 2022 | Co-Supervisor | Quantifying and Reducing Biases in Paleogenomic Research | Doctor of Philosophy | Doctorate | Full Time | Mr Adrien Oliva |
| 2017 - 2022 | Co-Supervisor | Exploring the Genetic History of the Ancient Near East through the Bronze and Iron Ages |
Doctor of Philosophy | Doctorate | Full Time | Mr Matthew Peter Williams |





Available For Media Comment.