
Dr Darren Wong
Future Making Fellow
Office of Sciences
College of Sciences
Eligible to supervise Masters and PhD - email supervisor to discuss availability.
Research Themes
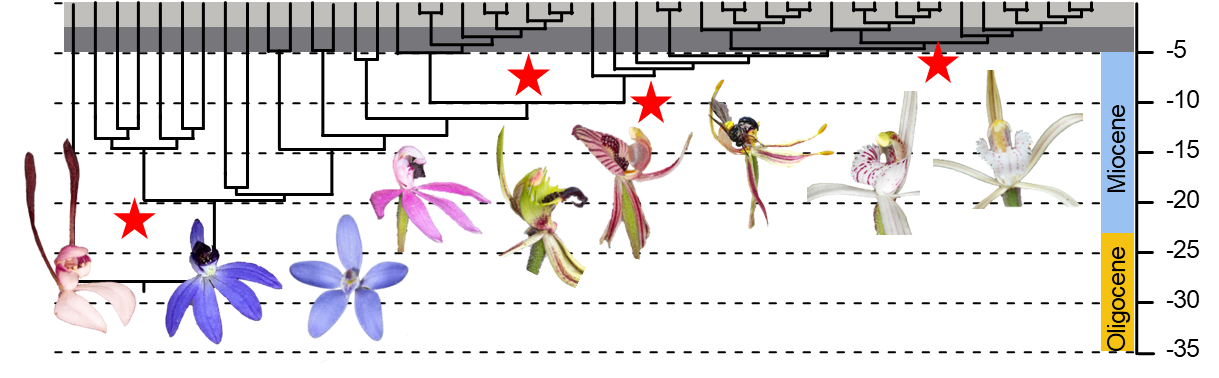
• Evolutionary adaptation in Australia's most charismatic plants
Revealing the genomic and metabolic innovations that drive rapid evolutionary change and unique floral traits in Australian orchids.

• Horticultural crop production & resilience
Dissecting the genetic basis of fruit ripening and the molecular responses that enable plants to withstand biotic stresses (pathogens, pests) and abiotic stresses (drought, cold, climate extremes).

• Emerging ethnobotanicals (Australian natives)
Leveraging modern analytical chemistry, genomics, and biotechnology with traditional knowledge to explore new applications of native plant chemistry.

Discovery to Delivery: From molecular mechanisms to real‑world impact
-
The Chemical Gap – Decoding adaptability
Mapping specialized metabolic pathways and identifying compounds underpinning stress tolerance and rapid evolution.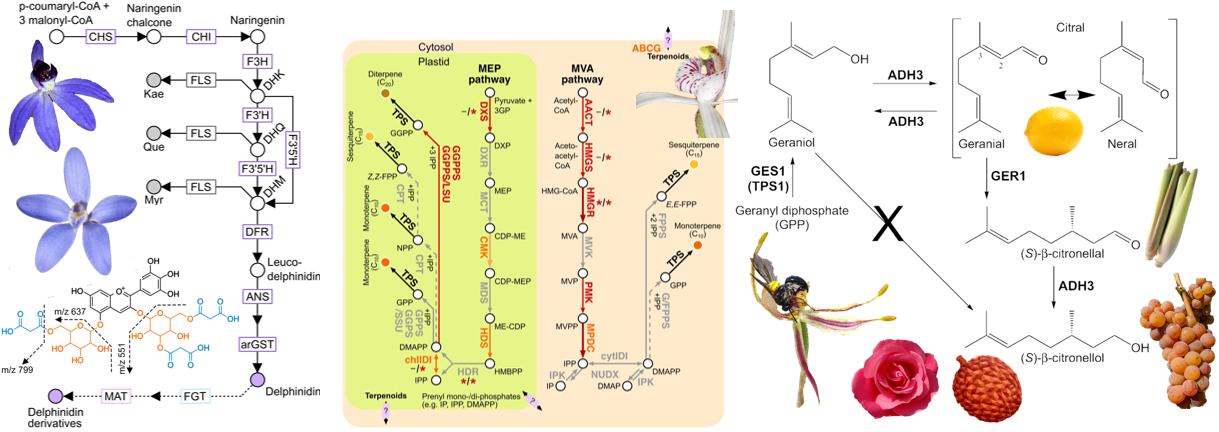
-
The Control Gap – Understanding genetic regulation
Uncovering the gene regulatory networks that control when and how these chemical pathways are switched on during development and environmental change.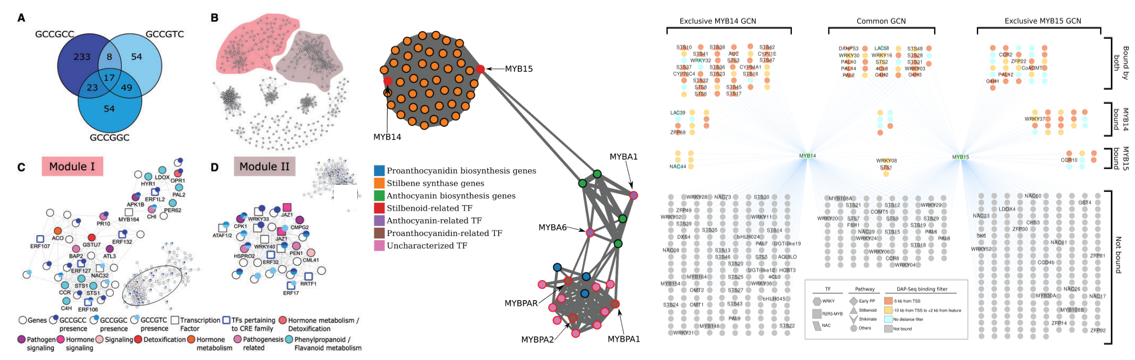
-
The Translation Gap – Scaling discovery
Using plant and microbial synthetic biology to sustainably produce high‑value compounds (e.g., unique fragrances and stabile pigments).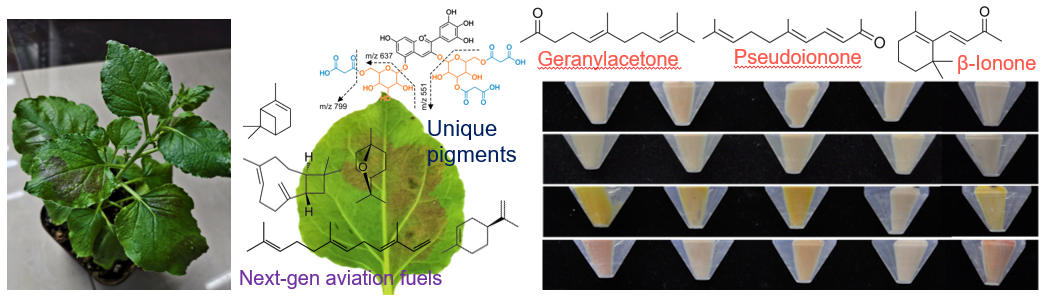
🌱 Join the team!
Exciting Honours, MSc, and PhD projects are available. Email your CV and research interest to darren.wong@adelaide.edu.au to discuss new opportunities.
My Google Scholar profile:
https://scholar.google.com/citations?user=rg8bEosAAAAJ&hl=en
Featured on the Cover (left to right): Plant Cell, Plant Journal, Journal of Experimental Botany, Biology of Plant Volatiles (2nd edition), Molecular Ecology

| Date | Position | Institution name |
|---|---|---|
| 2025 - ongoing | Future Making Fellow | University of Adelaide |
| 2025 - ongoing | Visiting fellow | Australian National University |
| 2023 - 2025 | Research fellow (level B) | Australian National University |
| 2019 - 2023 | DECRA fellow | Australian National University |
| 2016 - 2018 | Research fellow (level A) | Australian National University |
| 2014 - 2016 | Postdoctoral fellow | University of British Columbia |
| Date | Type | Title | Institution Name | Country | Amount |
|---|---|---|---|---|---|
| 2025 | Award | Best EMCR Presentation Award | TERPNET2025 | Australia | - |
| 2024 | Teaching Award | RSB Director's Prize in Honours | Australian National University | Australia | - |
| Date | Institution name | Country | Title |
|---|---|---|---|
| University of Adelaide | Australia | PhD | |
| University of Adelaide | Australia | B. Sc Hons (Plant Science) | |
| University of Adelaide | Australia | B. Sc (Genetics) |
| Year | Citation |
|---|---|
| 2026 | Wang, Z., Wang, Y., & Wong, D. C. J. (2026). Editorial: Surviving and thriving: how crops perceive and respond to temperature stress, volume II. Frontiers in Plant Science, 16, 3 pages. |
| 2026 | Hou, Y., Wong, D. C. J., Wang, L., Kang, Y., Zhou, H., Kafle, S., . . . Xin, H. (2026). A Hierarchical VvbHLH30-VvERF70-VvACS2 Module Orchestrates Ethylene Biosynthesis and Cold Adaptation in Grapevine.. Plant Biotechnol J, 25 pages. |
| 2025 | Wang, Z., Wang, Y., & Wong, D. C. J. (2025). Editorial: Surviving and thriving: how crops perceive and respond to temperature stress. Frontiers in Plant Science, 15, 1550257-1-1550257-4. Scopus5 WoS5 Europe PMC5 |
| 2025 | Zhou, F., Zhao, Y. N., Perkins, J., Xu, H., Pichersky, E., Peakall, R., & Wong, D. C. J. (2025). Fine-tuned terpene synthase gene expression, functional promiscuity, and subcellular localization: Implications for the evolution of complex floral volatile bouquet in Caladenia orchids. Plant and Cell Physiology, 66(4), 627-644. Scopus2 WoS2 Europe PMC2 |
| 2025 | Wong, D. C. J., Wang, Z., Perkins, J., Jin, X., Marsh, G. E., John, E. G., & Peakall, R. (2025). The road less taken: Dihydroflavonol 4-reductase inactivation and delphinidin anthocyanin loss underpins a natural intraspecific flower colour variation. Molecular Ecology, 34(15), e17334-1-e17334-21. Scopus7 WoS7 Europe PMC7 |
| 2024 | Hou, Y., Wong, D. C. J., Sun, X., Li, Q., Zhou, H., Meng, L., . . . Xin, H. (2024). VvbHLH036, a basic helix-loop-helix transcription factor regulates the cold tolerance of grapevine. Plant Physiology, 196(4), 2871-2889. Scopus16 WoS16 Europe PMC10 |
| 2024 | O’Donnell, R. P., Wong, D. C. J., Phillips, R. D., Peakall, R., & Linde, C. C. (2024). Discordance Down Under: Combining Phylogenomics and Fungal Symbioses to Detangle Difficult Nodes in a Diverse Tribe of Australian Terrestrial Orchids. Systematic Biology, 74(3), 434-452. Scopus3 WoS3 Europe PMC2 |
| 2023 | Wong, D. C. J., Pichersky, E., & Peakall, R. (2023). Many different flowers make a bouquet: Lessons from specialized metabolite diversity in plant–pollinator interactions. Current Opinion in Plant Biology, 73, 102332. Scopus14 Europe PMC12 |
| 2023 | Cardona-Medina, E., Santos, M., Nodari, R., Hornero-Méndez, D., Peris, A., Wong, D. C. J., . . . Rodríguez-Concepción, M. (2023). Accumulation of azafrin in the root apoplast of the medicinal plant Escobedia grandiflora might play a role in parasitism. Plants People Planet, 5(3), 354-367. |
| 2023 | Orduña, L., Santiago, A., Navarro-Payá, D., Zhang, C., Wong, D. C. J., & Matus, J. T. (2023). Aggregated gene co-expression networks predict transcription factor regulatory landscapes in grapevine. Journal of Experimental Botany, 74(21), 6522-6540. Scopus7 WoS6 Europe PMC12 |
| 2023 | Jin, X., Wang, Z., Li, X., Ai, Q., Wong, D. C. J., Zhang, F., . . . Si, H. (2023). Current perspectives of lncRNAs in abiotic and biotic stress tolerance in plants. Frontiers in Plant Science, 14. Scopus32 |
| 2023 | Zhang, C., Dai, Z., Ferrier, T., Orduña, L., Santiago, A., Peris, A., . . . Matus, J. T. (2023). MYB24 orchestrates terpene and flavonol metabolism as light responses to anthocyanin depletion in variegated grape berries. Plant Cell, 35(12), 4238-4265. Scopus42 WoS40 Europe PMC30 |
| 2023 | Hou, Y., Wong, D. C. J., Li, Q., Zhou, H., Zhu, Z., Gong, L., . . . Xin, H. (2023). Dissecting the effect of ethylene in the transcriptional regulation of chilling treatment in grapevine leaves. Plant Physiology and Biochemistry, 196, 1084-1097. Scopus21 Europe PMC12 |
| 2022 | Orduña, L., Li, M., Navarro-Payá, D., Zhang, C., Santiago, A., Romero, P., . . . Matus, J. T. (2022). Direct regulation of shikimate, early phenylpropanoid, and stilbenoid pathways by Subgroup 2 R2R3-MYBs in grapevine. Plant Journal, 110(2), 529-547. Scopus44 Europe PMC37 |
| 2022 | Sweetman, C., Waterman, C. D., Wong, D. C. J., Day, D. A., Jenkins, C. L. D., & Soole, K. L. (2022). Altering the balance between AOX1A and NDB2 expression affects a common set of transcripts in Arabidopsis. Frontiers in Plant Science, 13. Scopus7 |
| 2022 | Wong, D. C. J., & Peakall, R. (2022). Orchid Phylotranscriptomics: The Prospects of Repurposing Multi-Tissue Transcriptomes for Phylogenetic Analysis and Beyond. Frontiers in Plant Science, 13. Scopus22 |
| 2022 | Wong, D. C. J., Perkins, J., & Peakall, R. (2022). Anthocyanin and Flavonol Glycoside Metabolic Pathways Underpin Floral Color Mimicry and Contrast in a Sexually Deceptive Orchid. Frontiers in Plant Science, 13. Scopus21 |
| 2022 | Wong, D. C. J., Perkins, J., & Peakall, R. (2022). Conserved pigment pathways underpin the dark insectiform floral structures of sexually deceptive Chiloglottis (Orchidaceae). Frontiers in Plant Science, 13. Scopus8 |
| 2022 | Wang, Z., Wong, D. C. J., Chen, Z., Bai, W., Si, H., & Jin, X. (2022). Emerging Roles of Plant DNA-Binding With One Finger Transcription Factors in Various Hormone and Stress Signaling Pathways. Frontiers in Plant Science, 13. Scopus29 |
| 2021 | Burbidge, C. A., Ford, C. M., Melino, V. J., Wong, D. C. J., Jia, Y., Jenkins, C. L. D., . . . Sweetman, C. (2021). Biosynthesis and Cellular Functions of Tartaric Acid in Grapevines.. Front Plant Sci, 12, 1-22. Scopus78 WoS73 Europe PMC44 |
| 2021 | Peakall, R., Wong, D. C. J., Phillips, R. D., Ruibal, M., Eyles, R., Rodriguez-Delgado, C., & Linde, C. C. (2021). A multitiered sequence capture strategy spanning broad evolutionary scales: Application for phylogenetic and phylogeographic studies of orchids. Molecular Ecology Resources, 21(4), 1118-1140. Scopus17 Europe PMC13 |
| 2021 | Degu, A., Wong, D. C. J., Ciman, G. M., Lonardi, F., Mattivi, F., & Fait, A. (2021). Not just shrivelling: time-series profiling of the biochemical changes in Corvina (Vitis vinifera L.) berries subjected to post-harvest withering. Oeno One, 55(2), 115-129. Scopus7 |
| 2021 | Wang, Z., Wong, D. C. J., Wang, Y., Xu, G., Ren, C., Liu, Y., . . . Liang, Z. (2021). GRAS-domain transcription factor PAT1 regulates jasmonic acid biosynthesis in grape cold stress response. Plant Physiology, 186(3), 1660-1678. Scopus116 Europe PMC87 |
| 2020 | Livingston, S. J., Quilichini, T. D., Booth, J. K., Wong, D. C. J., Rensing, K. H., Laflamme-Yonkman, J., . . . Samuels, A. L. (2020). Cannabis glandular trichomes alter morphology and metabolite content during flower maturation. Plant Journal, 101(1), 37-56. Scopus242 Europe PMC155 |
| 2020 | Dimopoulos, N., Tindjau, R., Wong, D. C. J., Matzat, T., Haslam, T., Song, C., . . . Castellarin, S. D. (2020). Drought stress modulates cuticular wax composition of the grape berry. Journal of Experimental Botany, 71(10), 3126-3141. Scopus81 |
| 2020 | Perkins, M. L., Schuetz, M., Unda, F., Smith, R. A., Sibout, R., Hoffmann, N. J., . . . Samuels, L. (2020). Dwarfism of high-monolignol Arabidopsis plants is rescued by ectopic LACCASE overexpression. Plant Direct, 4(9). Scopus24 |
| 2020 | Kreynes, A. E., Yong, Z., Liu, X. M., Wong, D. C. J., Castellarin, S. D., & Ellis, B. E. (2020). Biological impacts of phosphomimic AtMYB75. Planta, 251(3), 60. Scopus12 Europe PMC9 |
| 2020 | Wong, D. C. J. (2020). Network aggregation improves gene function prediction of grapevine gene co-expression networks. Plant Molecular Biology, 103(4-5), 425-441. Scopus20 Europe PMC21 |
| 2020 | Azaman, S. N. A., Wong, D. C. J., Tan, S. W., Yusoff, F. M., Nagao, N., & Yeap, S. K. (2020). De novo transcriptome analysis of Chlorella sorokiniana: effect of glucose assimilation, and moderate light intensity. Scientific Reports, 10(1), 17331. Scopus31 Europe PMC14 |
| 2019 | Wong, D. C. J. (2019). Harnessing Integrated Omics Approaches for Plant Specialized Metabolism Research: New Insights into Shikonin Biosynthesis. Plant and Cell Physiology, 60(1), 4-6. Scopus12 Europe PMC9 |
| 2019 | Wong, D. C. J., Amarasinghe, R., Falara, V., Pichersky, E., & Peakall, R. (2019). Duplication and selection in β-ketoacyl-ACP synthase gene lineages in the sexually deceptive Chiloglottis (Orchidaceace). Annals of Botany, 123(6), 1053-1066. Scopus9 Europe PMC9 |
| 2019 | Sun, X., Zhang, L., Wong, D. C. J., Wang, Y., Zhu, Z., Xu, G., . . . Xin, H. (2019). The ethylene response factor VaERF092 from Amur grape regulates the transcription factor VaWRKY33, improving cold tolerance. Plant Journal, 99(5), 988-1002. Scopus131 Europe PMC83 |
| 2019 | Degu, A., Hochberg, U., Wong, D. C. J., Alberti, G., Lazarovitch, N., Peterlunger, E., . . . Fait, A. (2019). Swift metabolite changes and leaf shedding are milestones in the acclimation process of grapevine under prolonged water stress. BMC Plant Biology, 19(1), 69. Scopus47 Europe PMC22 |
| 2018 | Vannozzi, A., Wong, D. C. J., Höll, J., Hmmam, I., Matus, J. T., Bogs, J., . . . Lucchin, M. (2018). Combinatorial Regulation of Stilbene Synthase Genes by WRKY and MYB Transcription Factors in Grapevine (Vitis vinifera L.). Plant and Cell Physiology, 59(5), 1043-1059. Scopus128 Europe PMC90 |
| 2018 | Sun, X., Matus, J. T., Wong, D. C. J., Wang, Z., Chai, F., Zhang, L., . . . Xin, H. (2018). The GARP/MYB-related grape transcription factor AQUILO improves cold tolerance and promotes the accumulation of raffinose family oligosaccharides. Journal of Experimental Botany, 69(7), 1749-1764. Scopus88 Europe PMC63 |
| 2018 | Chen, Y., Grimplet, J., David, K., Castellarin, S. D., Terol, J., Wong, D. C. J., . . . Chervin, C. (2018). Ethylene receptors and related proteins in climacteric and non-climacteric fruits. Plant Science, 276, 63-72. Scopus135 Europe PMC61 |
| 2018 | Wong, D. C. J., Amarasinghe, R., Pichersky, E., & Peakall, R. (2018). Evidence for the involvement of fatty acid biosynthesis and degradation in the formation of insect sex pheromone-mimicking chiloglottones in sexually deceptive chiloglottis orchids. Frontiers in Plant Science, 9. Scopus9 |
| 2018 | Wong, D. C. J., Zhang, L., Merlin, I., Castellarin, S. D., & Gambetta, G. A. (2018). Structure and transcriptional regulation of the major intrinsic protein gene family in grapevine. BMC Genomics, 19(1), 248. Scopus29 Europe PMC18 |
| 2018 | Wong, D. C. J., Ariani, P., Castellarin, S., Polverari, A., & Vandelle, E. (2018). Co-expression network analysis and cis-regulatory element enrichment determine putative functions and regulatory mechanisms of grapevine ATL E3 ubiquitin ligases. Scientific Reports, 8(1), 3151. Scopus10 Europe PMC6 |
| 2018 | Vandelle, E., Vannozzi, A., Wong, D., Danzi, D., Digby, A. M., Dal Santo, S., & Astegno, A. (2018). Identification, characterization, and expression analysis of calmodulin and calmodulin-like genes in grapevine (Vitis vinifera) reveal likely roles in stress responses. Plant Physiology and Biochemistry, 129, 221-237. Scopus64 Europe PMC44 |
| 2017 | Xu, H., Bohman, B., Wong, D. C. J., Rodriguez-Delgado, C., Scaffidi, A., Flematti, G. R., . . . Peakall, R. (2017). Complex Sexual Deception in an Orchid Is Achieved by Co-opting Two Independent Biosynthetic Pathways for Pollinator Attraction. Current Biology, 27(13), 1867-1877.e5. Scopus74 WoS72 Europe PMC57 |
| 2017 | Wong, D. C. J., & Matus, J. T. (2017). Constructing integrated networks for identifying new secondary metabolic pathway regulators in grapevine: Recent applications and future opportunities. Frontiers in Plant Science, 8, 8 pages. Scopus58 WoS57 Europe PMC44 |
| 2017 | Wong, D. C. J., Pichersky, E., & Peakall, R. (2017). The biosynthesis of unusual floral volatiles and blends involved in orchid pollination by deception: Current progress and future prospects. Frontiers in Plant Science, 8. Scopus36 |
| 2017 | Wong, D. C. J., Amarasinghe, R., Rodriguez-Delgado, C., Eyles, R., Pichersky, E., & Peakall, R. (2017). Tissue-specific floral transcriptome analysis of the sexually deceptive orchid Chiloglottis trapeziformis provides insights into the biosynthesis and regulation of its unique UV-B dependent floral volatile, chiloglottone 1. Frontiers in Plant Science, 8. Scopus19 |
| 2017 | Wong, D. C. J., Lopez Gutierrez, R., Gambetta, G. A., & Castellarin, S. D. (2017). Genome-wide analysis of cis-regulatory element structure and discovery of motif-driven gene co-expression networks in grapevine. DNA Research, 24(3), 311-326. Scopus40 Europe PMC30 |
| 2017 | Savoi, S., Wong, D. C. J., Degu, A., Herrera, J. C., Bucchetti, B., Peterlunger, E., . . . Castellarin, S. D. (2017). Multi-omics and integrated network analyses reveal new insights into the systems relationships between metabolites, structural genes, and transcriptional regulators in developing grape berries (Vitis vinifera L.) exposed to water deficit. Frontiers in Plant Science, 8. Scopus112 |
| 2017 | Ariani, P., Vandelle, E., Wong, D., Giorgetti, A., Porceddu, A., Camiolo, S., & Polverari, A. (2017). Comprehensive workflow for the genome-wide identification and expression meta-analysis of the ATL E3 ubiquitin ligase gene family in grapevine. Journal of Visualized Experiments, 2017(130). Scopus5 Europe PMC3 |
| 2016 | Wong, D. C. J., Schlechter, R., Vannozzi, A., Höll, J., Hmmam, I., Bogs, J., . . . Matus, J. T. (2016). A systems-oriented analysis of the grapevine R2R3-MYB transcription factor family uncovers new insights into the regulation of stilbene accumulation. DNA Research, 23(5), 451-466. Scopus121 WoS108 Europe PMC95 |
| 2016 | Ariani, P., Regaiolo, A., Lovato, A., Giorgetti, A., Porceddu, A., Camiolo, S., . . . Polverari, A. (2016). Genome-wide characterisation and expression profile of the grapevine ATL ubiquitin ligase family reveal biotic and abiotic stress-responsive and development-related members. Scientific Reports, 6(1), 17 pages. Scopus31 WoS31 Europe PMC27 |
| 2016 | Loyola, R., Herrera, D., Mas, A., Wong, D. C. J., Höll, J., Cavallini, E., . . . Arce-Johnson, P. (2016). The photomorphogenic factors UV-B RECEPTOR 1, ELONGATED HYPOCOTYL 5, and HY5 HOMOLOGUE are part of the UV-B signalling pathway in grapevine and mediate flavonol accumulation in response to the environment. Journal of Experimental Botany, 67(18), 5429-5445. Scopus116 WoS116 Europe PMC88 |
| 2016 | Savoi, S., Wong, D. C. J., Arapitsas, P., Miculan, M., Bucchetti, B., Peterlunger, E., . . . Castellarin, S. D. (2016). Transcriptome and metabolite profiling reveals that prolonged drought modulates the phenylpropanoid and terpenoid pathway in white grapes (Vitis vinifera L.). BMC Plant Biology, 16(1), 17 pages. Scopus299 WoS280 Europe PMC172 |
| 2016 | Wong, D. C. J., Lopez Gutierrez, R., Dimopoulos, N., Gambetta, G. A., & Castellarin, S. D. (2016). Combined physiological, transcriptome, and cis-regulatory element analyses indicate that key aspects of ripening, metabolism, and transcriptional program in grapes (Vitis vinifera L.) are differentially modulated accordingly to fruit size. BMC Genomics, 17(1), 22 pages. Scopus59 WoS55 Europe PMC40 |
| 2015 | Jia, Y., Wong, D., Sweetman, C., Bruning, J., & Ford, C. (2015). New insights into the evolutionary history of plant sorbitol dehydrogenase. BMC Plant Biology, 15(1), 101-1-101-23. Scopus35 WoS33 Europe PMC22 |
| 2014 | Wong, D., Sweetman, C., & Ford, C. (2014). Annotation of gene function in citrus using gene expression information and co-expression networks. BMC Plant Biology, 14(1), 186-1-186-17. Scopus43 WoS40 Europe PMC32 |
| 2013 | Wong, D., Sweetman, C., Drew, D., & Ford, C. (2013). VTCdb: a gene co-expression database for the crop species Vitis vinifera (grapevine). BMC Genomics, 14(1), 1-17. Scopus56 WoS52 Europe PMC48 |
| 2012 | Sweetman, C., Wong, D., Ford, C., & Drew, D. (2012). Transcriptome analysis at four developmental stages of grape berry (Vitis vinifera cv. Shiraz) provides insights into regulated and coordinated gene expression. BMC Genomics, 12(691), 1-25. Scopus145 WoS134 Europe PMC92 |
| 2012 | Wong, D. J. C., Ismail, I. A., Godfrey, D., & Able, A. J. (2012). Death by toxin net blotch disease of barley. Microbiology Australia, 33(1), 34-35. |
| Year | Citation |
|---|---|
| 2020 | Peakall, R., Wong, D., Bohman, B., Flematti, G., & Pichersky, E. (2020). Chapter 15 Floral Volatiles for Pollinator Attraction and Speciation in Sexually Deceptive Orchids. In E. Pichersky, & N. Dudareva (Eds.), Biology of Plant Volatiles. CRC Press. DOI |
| 2020 | Wong, D., Peakall, R., & Pichersky, E. (2020). Chapter 6 The Role of Transcriptome Analysis in Shaping the Discovery of Plant Volatile Genes. In E. Pichersky, & N. Dudareva (Eds.), Biology of Plant Volatiles. CRC Press. |
| 2019 | Matus, J. T., Ruggieri, V., Romero, F. J., Moretto, M., & Wong, D. C. J. (2019). Status and Prospects of Systems Biology in Grapevine Research. In Compendium of Plant Genomes (pp. 137-166). Springer International Publishing. DOI |
| 2019 | Falchi, R., Wong, D. C. J., Yan, Y., Savoi, S., Gambetta, G. A., & Castellarin, S. D. (2019). The Genomics of Grape Berry Ripening. In Compendium of Plant Genomes (pp. 247-274). Springer International Publishing. DOI |
Cat 1 - Australian Competitive Grants - Commonwealth Funder
Title: Adaptation, evolution and conservation of Australia's diverse orchids (2026)
Description: This project uses comparative genomics to decode the evolutionary history of key orchid lineages. By translating these genetic insights into practical molecular tools, the work will advance species identification and conservation while fostering greater biotechnological innovation and environmental resilience.
Funding Scheme: ARC Discovery Project (DP)
Investigators: Peakall R (Lead), Wong DCJ (co-lead), Schwessinger B, Philipp S, Chagné D
Total Awarded Amount: AUD 903,000
Title: Genomic analysis of specificity and nutrient exchange in orchid-fungal interactions (2024)
Description: This project tackles a key unexplored aspect of molecular conservation biology of Australian orchids. Here we aim to identify genes that allow orchid mycorrhizal fungi to support orchid germination and growth.
Funding Scheme: Hermon Slade Foundation Grant
Investigators: Linde C (Lead), Peakall R, Schwessinger B, Wong DCJ
Total Awarded Amount: AUD 30,000
Title: The evolution of specialized orchid pollination and its reversibility (2021)
Description: This project aims to determine the changes in key floral volatile compounds underpinning pollination transitions, identify their molecular basis, and understand the ecological processes favouring reversals away from extreme specialization.
Funding Scheme: ARC Discovery Project (DP)
Total Awarded Amount: AUD 427,250
Title: Molecular systems biology of novel flower colour evolution (2019)
Description: This study will reveal how some plants evolve unique floral colour signals during speciation and lineage diversification in Australia.
Funding Scheme: ARC Discovery Early Career Researcher Award (DECRA)
Total Awarded Amount: AUD 392,000
Title: Identifying the molecular mechanisms of chickpea respiration and growth during stress using
integrated multi-omics analyses (2025)
Funding Scheme: South Australian Multiomics Framework Initiative (Bioplatforms Australia)
Investigators: Sweetman C (Lead), Hayes J (Co-lead), Sadras V, Cossani M, Lake L, Wong DCJ, Day D, Soole K, Jenkins C, Schleyer K
Other Funding:
Title: University of Adelaide Future Making Fellowship
Total Awarded Amount: AUD 400,000
Title: RSB Small Research Equipment Scheme
Description: Thermo/Dionex Diode Array Detector (DAD) add-on for Thermo Orbitrap QExactive Plus
Funding Scheme: RSB Small Research Equipment Scheme Funding Category
Total Awarded Amount: AUD 24,000
Title: Adelaide Graduate Research Award
Funding Scheme: Adelaide Graduate Research Award
Total Awarded Amount: AUD 90,000
ANU (2018-2023)
Advanced Research Techniques (BIOL8702): Led the curriculum redesign (2018–2019) and developed and delivered the high-impact Genomics and Next-Generation Sequencing (NGS) module (2018–2023), which accounts for 20% of the course assessment.
| Date | Role | Research Topic | Location | Program | Supervision Type | Student Load | Student Name |
|---|---|---|---|---|---|---|---|
| 2023 - 2024 | Principal Supervisor | Exploring the genetic mechanisms underpinning floral pigmentation variation in Australian Caladenia orchids | Australian National University | - | Honours | Full Time | Emma J |
| 2023 - 2023 | Principal Supervisor | Exploring the genomic basis underpinning the loss of floral pigmentation in the orchid Glossodia major | Australian National University | Advanced Studies Course (ASC) | Other | Part Time | Emma J |
| 2022 - 2023 | Principal Supervisor | Differentially expressed anthocyanin pathway genes in Caladenia carnea | Australian National University | Summer Research Internship | Other | Full Time | You J |
| 2021 - 2022 | Principal Supervisor | Genetic Mechanisms Underlying Changes in Flower Colour of Glossodia major | Australian National University | Summer Research Internship Program | Other | Full Time | Grace M |
| 2021 - ongoing | Co-Supervisor | Chemical Ecology and Pollination of the Eastern Australian Underground Orchids (Rhizanthella spp.) | Australian National University | - | Doctorate | Full Time | James P |
| Date | Topic | Location | Name |
|---|---|---|---|
| 2025 - ongoing | Evolution of Fruit Scent in Madagascar Figs | German Centre for Integrative Biodiversity Research (iDiv) | Linh N |
| 2022 - ongoing | Understanding Australian Orchids & Their Mycorrhizal Fungi – Macro to Microevolutionary Perspectives | Australian National University | Ryan P |
| Date | Role | Editorial Board Name | Institution | Country |
|---|---|---|---|---|
| 2023 - ongoing | Associate Editor | Frontiers in Plant Science | Frontiers Media SA | Switzerland |
| Date | Engagement Type | Partner Name |
|---|---|---|
| 2017 - 2020 | Consultant | BioCan Technologies Inc. |




